Evaluation data
Datasets
Public metadata are shown here; raw counts, cell barcodes, model weights, and unpublished result matrices remain outside the site.
Datasets
53
27 cancer and 26 development cohorts
Study tracks
11
7 comparative, 4 robustness and efficiency
Metrics
20
clustering, DRE, and LSE families
Primary latent
10D
shared dimensionality for learned methods
Inventory
Dataset identifiers
Cancer
Primary tumors, blood malignancies, metastasis samples, and cancer immune contexts.
| Subcategory | Count | Dataset identifiers |
|---|---|---|
Skin carcinoma Basal and squamous cell carcinoma cohorts. | 2 | GSE123813_bccHmCancerGSE123813_sccHmCancer |
Breast cancer Epithelial, stromal, primary, and metastatic breast cancer datasets. | 6 | GSE155109_bcECHmCancerGSE155109_bcStromaHmCancerGSE225600_breast_CancerHmGSE262288_breastMetasisHmCancerGSE168181_BreastHmCancerGSE228499_breastHmCancer |
Liver and metastasis Liver cancer, liver metastasis, and hepatoblastoma-related cohorts. | 4 | GSE98638_TcellLiverHmCancerGSE138709_LiverCancerGSE225857_liverColonMetasisHmCancerGSE283205_hepatoblastomaCancer |
Blood and lymphoid malignancy AML, ALL, multiple myeloma, and lymphoma contexts. | 5 | GSE132509_acutelymluekPBMCHmCancerGSE148218_bmALLHmCancerGSE222369_NKsLymphomaHmCancerGSE235787_bcellsALLHmCancerGSE124310_MMHmCancer |
GI tract Gastric, stomach, and colorectal adenocarcinoma contexts. | 3 | GSE183904_GastricHmCancerGSE149655_CAHmCancerGSE163558_stomachHmCancer |
Lung adenocarcinoma Two lung adenocarcinoma cohorts. | 2 | GSE123902_LungAdreHmCancerGSE189357_lungAdreHmCancer |
Brain metastasis Liver and triple-negative breast cancer brain-metastasis cohorts. | 2 | GSE143423_lbm_CancerBrainHmGSE143423_tnbc_CancerBrainHm |
Merkel cell carcinoma PBMC and tumor sampling contexts for Merkel cell carcinoma. | 2 | GSE117988_MCCPBMCCancerGSE117988_MCCTumorCancer |
T-cell cancers T-cell cancer immune-state dataset. | 1 | GSE222002_TcellsHmCancer |
Development
Hematopoietic, neural, embryonic, organ-development, disease-model, and atlas-scale systems.
| Subcategory | Count | Dataset identifiers |
|---|---|---|
Hematopoiesis CD34+ progenitors, HSC aging, bone marrow niche, and immune differentiation. | 8 | settyhematobm_GSE120446GSE253355_bmNicheHmGSE226131_HSCMmAgedGSE165844_LSKMmBatchGSE120505_bloodAgedifnHSPC_GSE226824 |
Neural development Dentate gyrus, spinal cord, retina, and astrocyte lineage contexts. | 4 | dentateGSE167597_spineMmGSE165784_RetinaHmDevGSE189070_astrocytesSCIMmDev |
Embryonic and stem-cell systems hESC time series, hESC-HSPC differentiation, and endoderm states. | 4 | GSE148215_hESCHSPCD8HmGSE192857_hESCHmTimeshESC_GSE144024endo |
Organ development Lung, pituitary, progastrin, urinary, and tooth development systems. | 6 | lungGSE130148_LungHmDevGSE142653pitHmDevGSE145929_ProgastinMmDevGSE145929_UrineMmDevGSE275119_TeethMmDev |
Disease models Inflammatory response and Alzheimer disease model contexts. | 2 | GSE115571_LPSMmDevGSE213740_ADHm |
Atlas references PanSci muscle and T-cell atlas-scale references. | 2 | GSE247719_PanSci_05_Muscle_adataGSE247719_PanSci_T_cell_adata |
Shared preprocessing
- Library-size normalization to 10,000 counts per cell
- log1p transform and selection of 2,000 highly variable genes
- Subsample to at most 3,000 cells with seed 42
- Leiden clustering at resolution 1.0 as the unsupervised reference partition
- Identical 15-nearest-neighbor graph and train-validation split for compared methods
Reference figure
Manuscript taxonomy figure
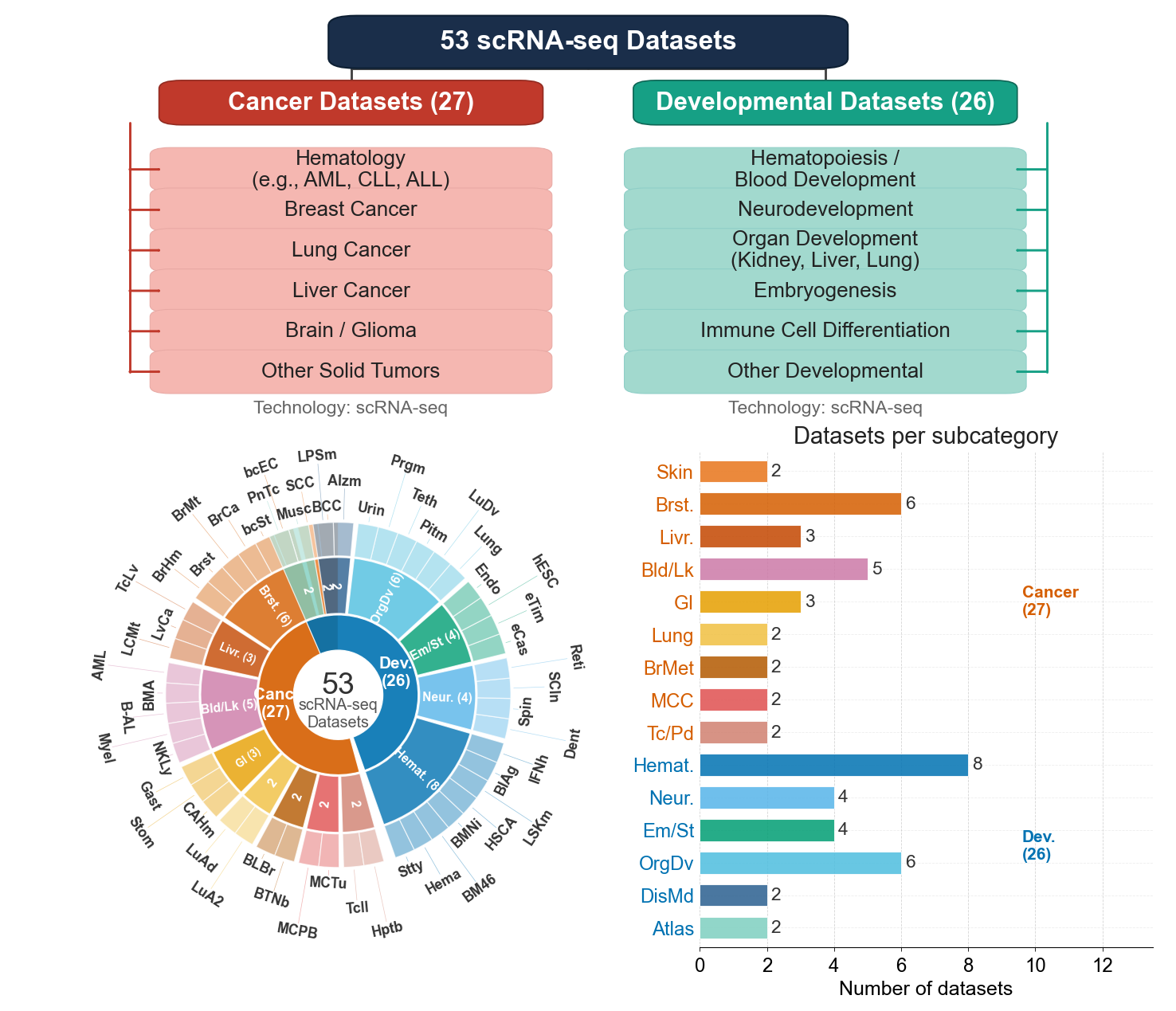
Acquisition
Datasets are obtained from their original publications or public archive
records. Public examples such as paul15 and
pbmc3k_processed are available through
standard Scanpy loaders; larger benchmark cohorts are resolved by the
experiment runners from local dataset directories.
Confidentiality note
This site does not host raw data, cell barcodes, sample identifiers, model weights, or unpublished result matrices. It documents the metadata needed to understand the evaluation design.
Continue